13 Soil Microbiomes
Soil Microbiomes
The terrestrial landscape on Earth harbors countless microbes that have major responsibilities in shaping their surrounding environment. One gram of soil can contain up to 10 billion microorganisms and consist of thousands of different species. Each ecosystem has unique soil properties that cultivate a diverse array of microbial communities, which are primarily composed of bacteria, however archaea, protists, fungi, viruses, and other microscopic organisms can be found in varying abundances too. The composition and health of the surrounding environment and its macro inhabitants are dependent on their soil borne microbial partners, and a disturbance in the balance of the soil microbiome can have far-reaching effects (Omotayo and Babalola, 2021).
Environmental and agricultural sustainability is important in a number of regards, especially maintaining biodiversity, plant and animal health, ecosystem homeostasis, and even human health. Anthropogenic effects on the climate and environment have become abundantly apparent, and it is imperative to adjust mindsets to become more environmentally conscious, not only to preserve aspects of nature, but to accommodate an increasing human population. The soil is foundational to agricultural practices, and therefore food production, and so a better understanding of various soil microbiomes could serve to benefit future applications and practices (Tosi et al., 2020). One possibility, is to become less reliant on synthetic nitrogen fertilizers for soil, and utilize symbiotic nitrogen-fixing bacteria as sources for plant nitrogen.
Microbe to Microbiome: A Paradigm Shift in the Application of Microorganisms for Sustainable Agriculture
Article by Ray et al., 2020 licensed under the terms of the Creative Commons Attribution License (CC BY).
Light, water and healthy soil are three essential natural resources required for agricultural productivity. Industrialization of agriculture has resulted in intensification of cropping practices using enormous amounts of chemical pesticides and fertilizers that damage these natural resources. Therefore, there is a need to embrace agriculture practices that do not depend on greater use of fertilizers and water to meet the growing demand of global food requirements. Plants and soil harbor millions of microorganisms, which collectively form a microbial community known as the microbiome. An effective microbiome can offer benefits to its host, including plant growth promotion, nutrient use efficiency, and control of pests and phytopathogens. Therefore, there is an immediate need to bring functional potential of plant-associated microbiome and its innovation into crop production. In addition to that, new scientific methodologies that can track the nutrient flux through the plant, its resident microbiome and surrounding soil, will offer new opportunities for the design of more efficient microbial consortia design. It is now increasingly acknowledged that the diversity of a microbial inoculum is as important as its plant growth promoting ability. Not surprisingly, outcomes from such plant and soil microbiome studies have resulted in a paradigm shift away from single, specific soil microbes to a more holistic microbiome approach for enhancing crop productivity and the restoration of soil health. Herein, we have reviewed this paradigm shift and discussed various aspects of benign microbiome-based approaches for sustainable agriculture.
Introduction
The health of soil plays an essential role in the ability of plants to produce food, fuel, and fiber for a growing world population. To keep pace, the total area of cultivated land worldwide has increased over 500% in the last five decades with a 700% increase in fertilizer use and a several-fold increase in pesticide use (Banerjee et al., 2019). In addition to being the world’s largest agricultural producers and exporters, the EU, Brazil, United States, and China also are some of the world’s largest pesticide users – each using 827 million, 831 million, 1.2 billion, and 3.9 billion pounds of pesticides, respectively, in 2016 (Donley, 2019). However, these numbers are not sustainable from either a supply-chain or environmental perspective. Thus, because natural resources are limited and their overuse pollutes the environment, the continued use of fertilizers and water to meet the demand of future global food requirements is not sustainable. Of relevance here is that agricultural intensification with high resource use and low crop diversity can negatively affect soil- and plant-associated microbiota (the so-called “phytobiome”) with subsequent impacts on critical ecosystem services (Matson et al., 1997).
There is growing evidence that aboveground plant diversity supports belowground microbial biodiversity, primarily through root exudation and rhizo-deposition (Bais et al., 2006; Eisenhauer et al., 2017; Morella et al., 2020). These more simple carbohydrates released into the soil primarily feed bacteria (Gunina and Kuzyakov, 2015) and are the most abundant near the root surface and diffuse along a gradient as distance from the root increases (Gao et al., 2011). The microbial composition is more abundant and complex in the rhizosphere, the narrow zone surrounding plant roots, with up to 109 cells per gram in typical rhizospheric soil, comprising up to 106 taxa (Lakshmanan et al., 2017). The more complex carbohydrates (e.g., lignin, cellulose) are largely degraded by decomposer fungi that break down these recalcitrant compounds into forms that can be used by other microbes. This conversion is largely decoupled from conventional agricultural practices, wherein the organic matter content is often lost to the system (Craven and Ray, 2019), and the carbon flux is at least partially unregulated in this regard. Again, defining nutrient fluxes with techniques like Stable Isotope Labeling (SIP) holds great potential to define and construct resilient, functioning and beneficial microbiomes that can contribute to future holistic agriculture. Thus, applying an efficient and diverse soil microbiome backed by these new technologies can facilitate and promote sustainable agriculture and can effectively contribute to meet the triple requirements of economic, social and environmental sustainability (Ray and Craven, 2016).
Historically, microorganisms that promote plant growth and nutrient acquisition have been used largely as single strains in agriculture to offset such fertilizer inputs as nitrogen and phosphorous. However, studies of natural populations suggest that groups of microbes with distinct function niches play pivotal roles in adhering and desorbing inorganic nutrients to physical surfaces, as well as breaking down organic residues and incorporating them into the soil (Lakshmanan et al., 2014; Finkel et al., 2017; Kumar and Dubey, 2020). Conceptually, such observations support the idea of the microbiome as a second genome or an extended genome of the plant (Vandenkoornhuyse et al., 2015). It is now evident that improving plant performance in a sustainable manner is beyond the binary interaction between a specific microbe or a consortium of beneficial microbes and a targeted host plant. This is a much more complex set of interactions than previously thought that requires modeling for improving predictable outcomes. In this review, we will highlight the current state of the art for the incorporation of specific plant growth-promoting microorganisms and discuss the principles and management practices for next-generation, microbiome-based approaches for sustainable agriculture.
Application of Beneficial Microbes in Sustainable Agriculture: Past, Present and Future
Since the early 1800s, the United States Department of Agriculture has recommended the use of certain rhizobacteria to improve nitrogen fertility in leguminous crops (Schneider, 1892). Since that time, a great deal of research has been conducted on this relationship between legumes and these bacteria, now termed rhizobia, that inhabit unique structures, the nodules, that form on the roots. Rhizobia infecting these nodules are now capable of “biological nitrogen fixation,” whereby di-nitrogen is fixed into forms that can be used by the plant. Symbiotically, the bacteria trade these nitrogenous compounds to the host plant in exchange for photosynthetically derived carbon. Despite these limited applications, much remains to be learned regarding both the functional and taxonomic diversity of these symbiotic bacteria and their host plants, the role they play in the global nitrogen cycle, and ultimately, how they can best be harnessed for improving plant productivity. This is particularly true for marginal lands that are not suited for row crop production but will need to be incorporated into global food and forage production approaches moving forward. Further, such degraded lands must but regenerated with the goal of restoring soil health and productivity. Any successful endeavor in this regard must include a characterization of the soil microbiome, both taxonomically and functionally. Attempts currently are underway to fix nitrogen in such non-legumes as wheat, corn and other staple crops that produce the bulk of human food by engineering symbiotic relationships using synthetic biology approaches (Rogers and Oldroyd, 2014; Ryu et al., 2020). Such approaches would significantly impact global food supplies, and may function adequately to reduce the arable land required to meet productivity goals.
Plant growth-promoting microbes not only play critical and diverse roles in growth promotion per se, but also in improving various aspects of plant resilience against a wide array of biotic and abiotic stresses (Arnold et al., 2003; Sun et al., 2010; Agler et al., 2016; Azad and Kaminskyj, 2016; Singh, 2016; Oleńska et al., 2020; Rai et al., 2020). In this context, researchers globally have worked over the last several decades on plant growth-promoting microorganisms, such as root-associated mycorrhizal fungi, across a broad range of crops and encompassing a wide range of agro-climatic conditions. For perspective, Brundrett and Tedersoo (2018) recently reviewed 135 years of mycorrhizal research and reported that merely 8% of the vascular plants are non-mycorrhizal, suggesting that plant families associating with mycorrhizae have been very successful over the evolution of the plant kingdom.
Traditionally, agricultural application of beneficial microorganisms involves a few types of well-characterized microbes, such as mycorrhizal fungi or rhizobia bacteria, for which the mechanisms underlying the plant growth promotion effects are well understood. Further, most of these studies focused solely on the ability of the applied microorganisms to facilitate such specific plant growth-promoting traits as phosphate solubilization, nitrogen fixation, ACC deaminase production (Sarkar et al., 2018), siderophore production, biofilm formation, plant hormone production, biotic, and abiotic stress tolerance or resistance, among others (Weyens et al., 2009; Bhattacharyya and Jha, 2012; Singh et al., 2019). While these beneficial microorganisms can impart considerable benefits to plant growth and fitness, they are typically documented in simple, one-on-one studies, often conducted in sterile soils in greenhouse conditions. As a consequence, the effects found in such simplified conditions often fail to translate to more complex field situations (Chutia et al., 2007; Nicot et al., 2011; Parnell et al., 2016). Soil in field plots have more complex microbial environments that are presumably adapted to the local eco-environment.
In recent years, next-generation sequencing has revolutionized our understanding of microbial community composition and function, and together with improved culturing methodologies has greatly facilitated the use of biologicals in the field (Schweitzer et al., 2008; Panke-Buisse et al., 2014; Mueller and Sachs, 2015). Specifically, metagenomics-based approaches have uncovered vast, previously unrecognized populations of microbes that may have new or enhanced properties that could be used for agriculture, bioremediation, and human health. For example, comparative analyses of rhizosphere metagenomes from resistant and susceptible tomato plants enabled the identification and assembly of a flavobacterial genome that was far more abundant in the resistant plant rhizosphere microbiome than in that of the susceptible plants. Such findings certainly reveal a role for native microbiota in protecting plants from phytopathogens, and pave a way forward for the development of probiotics to ameliorate plant diseases akin to human health (Kwak et al., 2018). In another study, a 16S rRNA gene amplicon sequencing analysis of maize root microbiome led to the identification of bacteria that promote growth under low temperature conditions (Beirinckx et al., 2020). Additionally, principles of consortium design that rely on cross-talk, cross-feeding and/or substrate channeling between different microorganisms offer new opportunities for “intelligent” consortia design (Calvo et al., 2014; Vorholt et al., 2017; Paredes et al., 2018). We propose that the manipulation of the plant microbiome holds tremendous potential for agricultural improvement (Table 1). Through recent years of research, it is elucidated how microbes worked in nature before, and how decades of chemical fertilizer use have silenced their ability to improve plant fitness and soil health. Therefore, designing a microbial consortium that carefully weighs and evaluates the relationship between inoculants and the resident microbiome would substantially improve the plant growth-promoting potential and resilience of agricultural biologicals to boost plant growth. In this review, we will discuss the key considerations that would improve the likelihood of microbial products to improve crop yield, decrease disease severity and/or ameliorate abiotic stress response. Further, it is likely that such considerations would reduce the inconsistency between the performances of beneficial microbes from controlled greenhouse conditions and more natural environments.

Microbes for Plant Growth Promotion: A Reductionist Approach
Sustainable agriculture primarily focuses on reducing the dependency of plants on chemical fertilizers and improving their ability to grow on marginal soil types. For such purposes, individual microorganisms for plant growth-promotion have largely focused on those that facilitate growth and development by enhancing acquisition of nutrient resources from the environment, including fixed nitrogen, iron and phosphate, or modulating growth by altering plant hormone levels (Figure 1) (Hayat et al., 2010). Another approach aimed at reducing yield losses to disease relies on microbes that decrease or prevent the deleterious effects of plant pathogens by several different mechanisms (Glick, 2012), i.e., by acting as a biocontrol agent. Microbe-based plant growth-promoting products, more popularly marketed as biofertilizer, has been commercially available in many countries since the 1950s (Timmusk et al., 2017). Application of such plant growth-promoting microbes in agricultural context and more specifically as inoculants has been nicely reviewed by Souza et al. (2015). However, under certain cases, the results obtained in the laboratory could not be reproduced in the field primarily due to the presence of many crop species and crop varieties, variable environmental conditions between fields, (Timmusk et al., 2017; Saad et al., 2020), occasionally due to the low quality of the inocula, and their inability to compete with the indigenous population. In that context, it is important to consider the fact that there is always greater likelihood of success by introducing mixed cultures of compatible microorganisms, rather than single, pure cultures. This is simply because each strain in the multi-strain consortium can compete effectively with the indigenous rhizosphere population and enhance plant growth with its partners. For example, sequential inoculation of nitrogen fixing bacterium Azotobacter vinelandii, followed by plant growth-promoting root-endophytic fungus Serendipita indica demonstrated better growth in rice (Dabral et al., 2020). Dual inoculation of S. indica and Mycolicibacterium strains boosted the beneficial effects in tomato (del Barrio-Duque et al., 2019) and that of arbuscular mycorrhizal fungus with plant growth-promoting bacteria Bacillus subtilis demonstrated better growth in wheat (Yadav et al., 2020) as compared to the singly inoculated plants. There also are numerous other reports that showed two strains used in a consortium promoted plant growth in a more effective manner (Nadeem et al., 2013; Fatnassi et al., 2015; Priyadharsini and Muthukumar, 2016). Nevertheless, to unlock the full potential of soil microbes for such nutrient cycling as nitrogen or phosphorus and providing plant protection against biotic and abiotic stress microbiomes, it is necessary to develop strategies to comprehend the functional capabilities of soil microbial communities. Irrespective of the approach, persistence is the first and foremost principle underlying the design of a successful microbial consortium for conferring plant growth promotion. This is not surprising, as the survival and activity of microbes in any soil system face a monumental task of competing with the myriad of microbes naturally adapted to that same soil. Thus, in addition to establishment of a compatible interaction with the host, a successful microbial inoculant has to subsequently compete and persist in the context of indigenous microbes as well as local abiotic conditions (Finkel et al., 2017). It has been reported that bacterial inoculations can persist in soil up to 7 weeks, but whether this inoculum also can provide plant growth benefits is not clear (Schreiter et al., 2014). While persistence or resilience of any microbial inoculum is more dependent on biotic components of a specific soil type, their persistence can be improved by inoculating crops with consortia rather than single strains (Verbruggen et al., 2012; Nemergut et al., 2013). Thus, it can arguably be stated that the diversity of a microbial inoculum, in addition to its plant growth-promoting traits, is critical for enhancing productivity and longevity (Cordero and Polz, 2014).
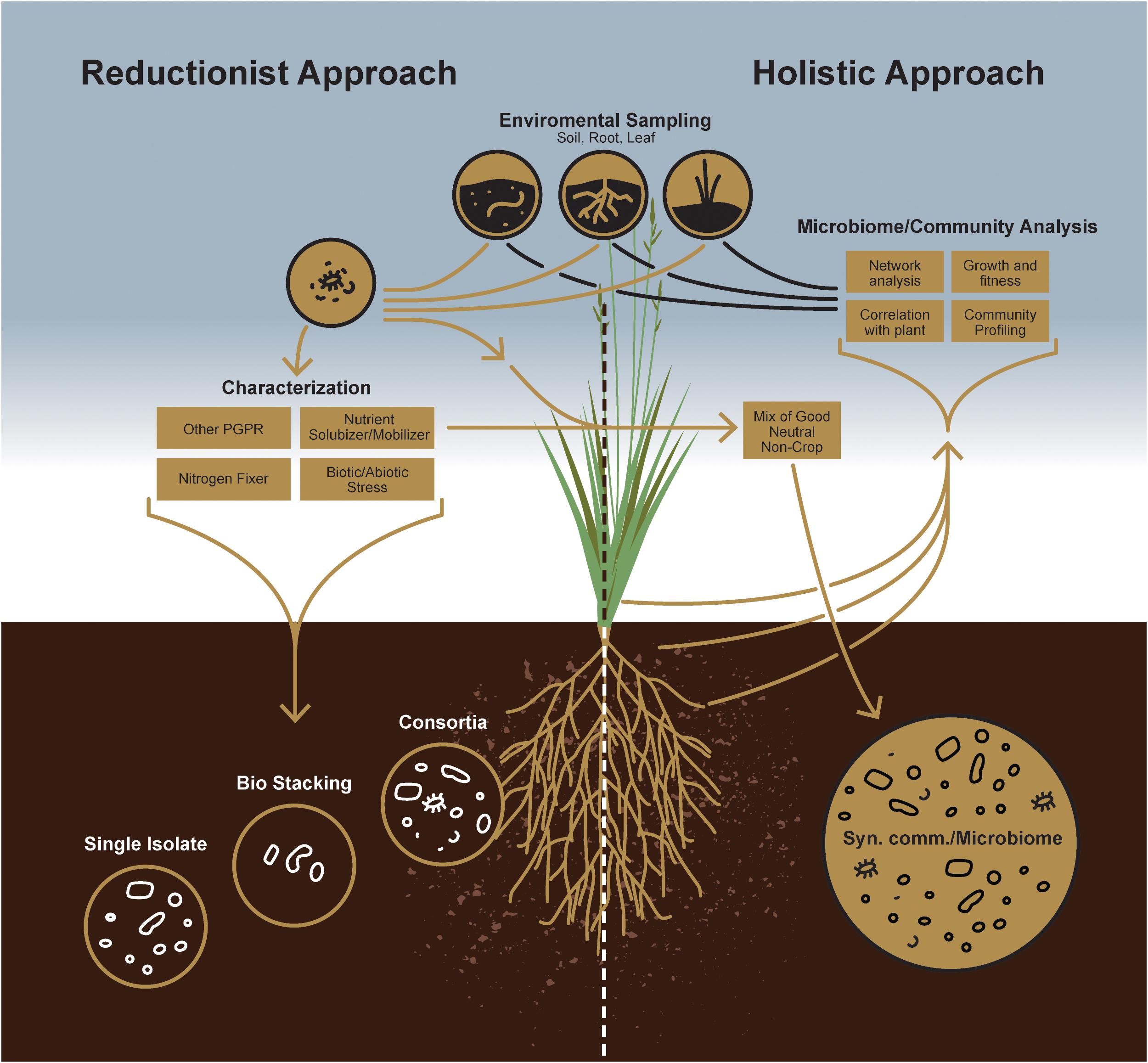
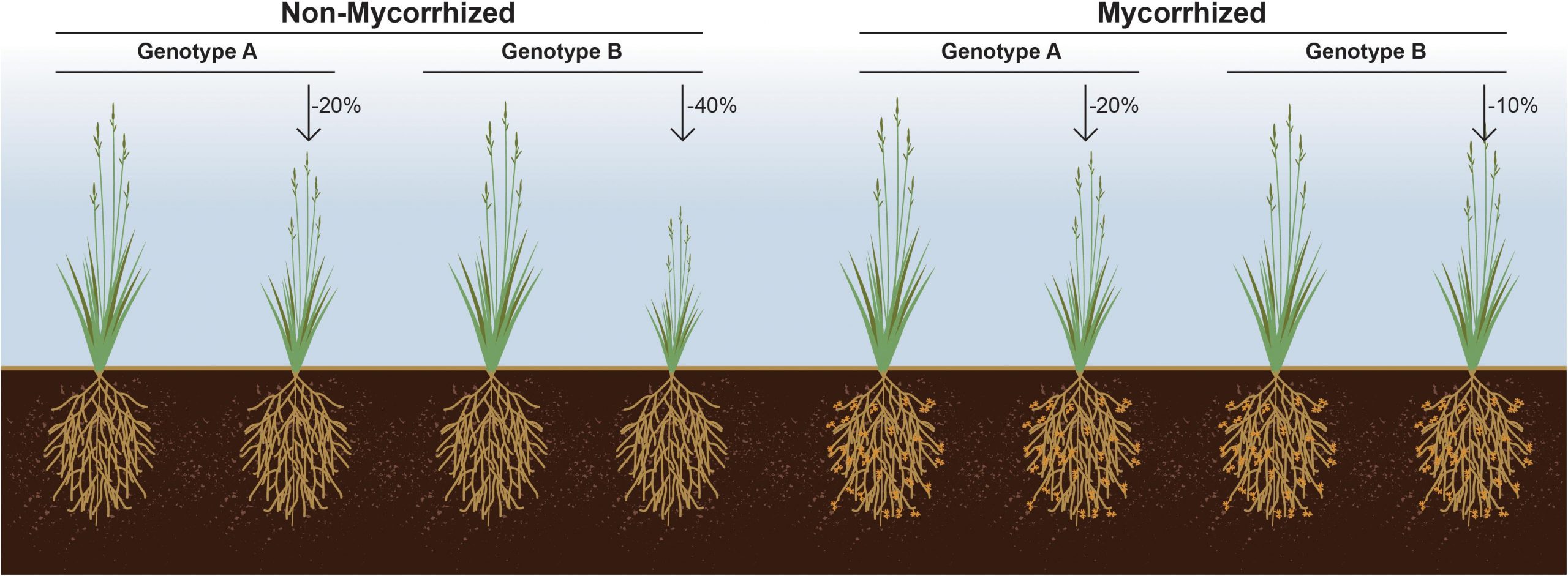
To improve the likelihood of success for such a management strategy, a priori knowledge of indigenous microbial populations competing with the introduced plant growth-promoting agent(s) is critical. While a reductionist approach can define the currency of individual plant-microbe interactions, the concepts of microbial community survival and functioning require, a more holistic, microbiome-based approach empowered by next-generation sequencing technology to study plant-microbe interactions at the community level (Figure 2). Indeed, this will enable researchers to design more robust, synthetic microbial consortia capable of reliably enhancing agricultural productivity.
Microbiomes for Plant Growth Promotion: The Holistic Approach
Soil is a vastly heterogeneous growth medium, providing a wide spectrum of ecological niches for microorganisms that enable diverse strains to coexist and form complex microbial communities. When the earliest plants extended their roots into primordial soils, they encountered a habitat already teeming with bacterial and fungal life (Bulgarelli et al., 2013; Kemen, 2014). Since that early time, plants have interacted with rhizosphere microbes, evolving strategies to forge beneficial alliances with some while keeping others at bay. Such early associations certainly had consequences on plant growth and development. Therefore, a more holistic approach is needed to understand better these microbes and the roles they play in the overall health of plant and soil (Figure 1). Again, recent advances in next-generation sequencing technology and the decreasing costs associated with that technology now allow us to evaluate how microbial populations fluctuate in both space and time or to identify core microbiomes that appear conserved among host genotypes or species (Sergaki et al., 2018). Thus, although culture-independent methods have contributed tremendously to our understanding of plant-associated fungal and bacterial community structures, the study of microbiome functions remains challenging because of the inherent noise of plant-associated microbial communities. It is now well known that there are core sets of microbes that, depending on the host, are recognized as keystone taxa that consistently associate with healthy plants (Banerjee et al., 2018). Consequently, researchers working with specific plant-microbe interactions have increasingly acknowledged the mitigating impact these larger microbial communities have on individual plant-microbe outcomes for plant growth promotion or fitness. Now, plant-associated fungal and bacterial stains from various plant species are being isolated, which will provide in the near future an inestimable resource for assembling taxonomically defined microbial communities with increasing complexity. Therefore, it is now imperative to take advantage of this knowledge to design consortia of microbes to maintain a sustainable rhizosphere community, with key functional properties that include plant protection, nutrient acquisition, and alleviating biotic and abiotic stress responses. From that perspective, synthetic community (SynCom) approaches can provide functional and mechanistic insights into how plants regulate their microbiomes (Figure 1). Not surprisingly, recent culture-independent analyses thus have paved the way for developing SynComs more often (Bodenhausen et al., 2014; Armanhi et al., 2018; Carlström et al., 2019).
Mycorrhizal fungi, at least the arbuscular type, were early symbiotic partners of most land plant species, improving nutritional conditions through soil exploration and pathogen resistance of host plants (Klironomos et al., 2000). In reward for the essential physiological services, they receive ca. 20% of net photosynthetic products from plants (HoÈgberg et al., 2001). Other mycorrhizal systems may have different nutritional benefits and costs, as has been proposed for the serendipitous system (Craven and Ray, 2019). Additionally, third-party partners can modulate the outcome of the tripartite interaction, such as the case of mycorrhizal helper bacteria (Frey-Klett et al., 2007), fungal endobacteria (Bonfante and Desirò, 2017; Bonfante et al., 2019) like Candidatus Moeniiplasma glomeromycotorum within the spores and hyphae of Glomeromycotina (Naito et al., 2017), Rhizobium radiobacter within Serendipita indica (Guo et al., 2017), and N2-fixing endobacteria Pseudomonas stutzeri inside basidiomycetes yeast endophyte Rhodotorula mucilaginosa (Paul et al., 2020). Hence, it is imperative to consider the composition and functioning of these microbe–microbe interactions to understand plant–microbiome associations in a holistic manner.
Principles and Management of Rhizosphere Microbiomes for Sustainable Agriculture
Competence and Resilience of the Rhizosphere Microbiome: Impact of Introduced Microbes on Native Microbiomes
In 1904, the German agronomist and plant physiologist Lorenz Hiltner coined the term rhizosphere (Hartmann et al., 2008) to describe the area around a plant root inhabited by a unique population of microorganisms. Since then, numerous studies have been undertaken to decipher the interplay between plants and rhizosphere microorganisms, encompassing a wide variety of plant growth-promoting bacteria, fungi, insects, protozoans, viruses, etc. (Marschner, 2012; McNear, 2013). The majority of these studies have traditionally followed a simple principle for maximizing successful host infection by pre-inoculation onto the targeted crop of choice to provide a competitive advantage for a desired microbe. Conceptually, this increases the relative abundance of a given beneficial microbe in the rhizosphere, at least temporarily, to achieve the desired benefit. Such studies typically take place in a controlled, artificial condition, such as a defined growth medium in a greenhouse, where competition from a native rhizosphere community is relatively low or non-existent. As mentioned above, this approach occasionally has failed once field application is attempted or the benefits are dramatically reduced in amplitude and/or endurance.
As an example, Lekberg and Helgason (2018) conducted a literature survey of research papers published on mycorrhizal functioning spanning a 30-year period (1987–2017). The most striking finding of this survey was that less than 5% of the work scientifically manipulated mycorrhizal abundance in the field. While we are not arguing the merit of greenhouse-based studies where the number of variables can be controlled and accounted for, yield gains in field conditions will continue to be modest with such an approach. Rhizosphere competence must be evaluated in a field situation if the true power of this approach is to be realized.
Over the last few decades, mycorrhiza-based bio-fertilizers containing one or several species of fungi were developed in forestry and agriculture (Jeffries and Rhodes, 1987; Baraza et al., 2016; Igiehon and Babalola, 2017). These inoculants are generally effective in plant growth promotion under controlled lab and greenhouse conditions. However, few targeted efforts have been made to measure interactions between the introduced microbe(s) and the native mycorrhizal community, let alone the more complex rhizosphere microbiome (Svenningsen et al., 2018; Turrini et al., 2018). To optimize outcomes from these interactions, targeted research must be undertaken to understand how such mycorrhiza-based biofertilizer integrate themselves within the context of the native microbiome.
Integration of Rhizosphere Microbiomes in Plant-Microbe-Nutrient Relationships
The soil microbial community often assists plants by weathering minerals from rock surfaces and degrading recalcitrant soil organic matter whereby soil microbes break down soluble and insoluble organic matter and convert it into inorganic, plant-available forms. Soil organic matter turnover is thus considered a net positive, as it liberates the nutrients locked up in organic matter. For this reason, conventional farming has always relied heavily on soil tillage, along with such other intensive agricultural practices as usage of inorganic fertilizers, herbicides and pesticides. However, it is already clear that such practices have negative consequences on the functional diversity of soil microbiomes. Long-term chemical fertilization has been shown to dramatically decrease the soil pH, which leads to a decrease in bacterial diversity and other changes in microbial community structure (Sun et al., 2015). This was well documented in the work of Kumar et al. (2017), who showed that long-term application of high doses of inorganic nitrogenous fertilizers severely reduces relative abundance, diversity and structure of diazotrophs, which play a key role in converting atmospheric N2 to plant-available ammonium.
As mentioned above, soil bacterial communities play a pivotal role in soil organic matter decomposition. In particular, soil carbon and nitrogen are critical factors for bacteria that rely on soil organic C and N decomposition to obtain energy (Chen et al., 2014; Wild et al., 2014; Tian et al., 2018). Further, different types of soil C selectively manipulate soil microbial community composition, resulting in changes in such belowground ecosystem functions as decomposition and nutrient transfer and creating feedbacks that may affect overall plant growth and productivity (Orwin et al., 2006). For example, bacteria belonging to the genera Chloroflexi, Nitrospirae, and Planctomycetes preferentially feed on recalcitrant organic C, whereas Proteobacteria and Bacteroidetes prefer labile organic C present in the soil (Nie et al., 2018). For this reason, amending the soil with such organic fertilizers as compost or manure contributes to higher microbial diversity and biomass compared to mineral-fertilized soils, which in turn positively impacts soil health (Schmid et al., 2018; Banerjee et al., 2019). Unfortunately, only a few agroecosystem experiments exist that compare organic and conventional management strategies over an extended period for evaluation of impact on soil health and restoration (Raupp et al., 2006; Khatoon et al., 2020). Hartmann et al. (2015) took a metagenomics approach to assess microbial diversity of soil in response to more than 20 years of continuous organic and conventional farming. Not surprisingly, they found that organic farming increased richness, decreased evenness, and shifted the structure of the soil microbiota when compared with conventionally managed soils under mineral fertilization (Hartman et al., 2018; Li et al., 2020b). There also are reports of significant alterations in the microbial community composition of both summer maize and winter wheat in response to increased nitrogen fertilization dose (Wang et al., 2018; Li et al., 2020a). Clearly, a better understanding of the interactions between the soil microbiome and conventional agricultural practices is crucial for the development of sustainable management of soil fertility and crop production.
Managing the Rhizosphere Microbiome to Induce Disease Suppression in Soil
Disease suppressive soils were originally defined by Baker and Cook (1974) as “soils in which the pathogen does not establish or persist, establishes but causes little or no damage, or establishes and causes disease for a while but thereafter the disease is less important, although the pathogen may persist in the soil.” Disease suppressive soils are the best examples of microbiome-mediated protection of plants against root infections by soil-borne pathogens. Such disease-suppressive soils have been described for various soil-borne pathogens, including fungi, bacteria, oomycetes, and nematodes (Mazzola, 2007; Kwak et al., 2018). To date, several microbial genera have been proposed as key players in disease suppressiveness of soils, but the complexity of the microbiome, as well as the underlying mechanisms and microbial traits, remain elusive for most disease suppressive soils (Toyota and Shirai, 2018).
Recently, Carrión et al. (2019) showed that upon pathogen invasion, members of the Chitinophagaceae and Flavobacteriaceae became enriched within the plant endosphere. They proposed that this bacterial population shift led to the induction of enzymatic activities associated with fungal cell-wall degradation, as well as secondary metabolite biosynthesis, all aimed at accelerating and augmenting the plant defense response(s). Although the disease suppressive abilities of certain soils can be at least partially attributed to their physico-chemical properties, the capacity of a soil to suppress disease progression is more often attributed to agri-management practices and crop rotation (Weller et al., 2002). In classic studies by Gerlagh (1968) and Shipton et al. (1973), the authors have shown soil to become disease suppressive after mono-culturing wheat over time. More recently, a comparative metatranscriptome analysis of wheat rhizosphere microbiome grown in fields suppressive and non-suppressive to the plant pathogen R. solani AG8 clearly revealed distinct dominant taxa in these two soil types. Additionally, suppressive samples showed greater expression of polyketide cyclase, terpenoid biosynthesis, and cold shock proteins (Hayden et al., 2018). While development of probiotics for the human gut microbiome has already been an established field of research, the use of probiotics that comprises naturally occurring bacterial antagonists and competitors that suppress pathogens has recently emerged as a promising strategy for disease suppression in soil. A study on application of probiotic consortia that comprised predefined Pseudomonas species reported suppression of the bacterial plant pathogen Ralstonia solanacearum in the tomato rhizosphere microbiome (Hu et al., 2016). In another study, amendment of Metarhizium, an insect-pathogenic fungus that is commonly employed as biological control agents against crop pests, in the rhizosphere of common bean (Phaseolus vulgaris) significantly increased the relative abundance of plant growth promoting such taxa as Bradyrhizobium, Flavobacterium, Chaetomium, and Trichoderma while suppressing the root rot disease symptoms Fusarium solani (Barelli et al., 2020). Soil suppressive properties are mostly derived from the biological functions of soils. Therefore, elucidation of microbial functions in suppressive soils by a next-generation sequencing approach will facilitate the development of effective, consistent and durable disease management tools.
Impact of Agriculture Management Practices on the Soil Microbiome
One important context for plant-microbe interactions is soil structure, as it can vary greatly depending on land-use history, plant species composition and successional stage (Erktan et al., 2016). Besides playing pivotal roles in soil organic matter decomposition, carbon cycling, nutrient mobilization, etc., saprotrophic fungi also are involved in creating soil structure through the secretion of extracellular compounds and physical binding of soil via hyphal networks (Bergmann et al., 2016). Interestingly, studies on the impact of tillage on the soil fungal communities have shown mixed results. Reports in no-till systems have varied from increased ratios of fungal to bacterial biomass (Acosta-Martínez et al., 2010) to decreased ratios (Mbuthia et al., 2015), as well as no change at all (Mathew et al., 2012). More recent studies have shown that soil fungal communities are negatively impacted by tillage, as they typically would be responsible for degrading crop residue left on the surface with no-till (Yin et al., 2017). More specifically, soil bacterial communities were primarily found to be structured by tillage, whereas soil fungal communities responded mainly to management type with additional effects by tillage (Hartman et al., 2018). Additionally, it is acknowledged that organically managed systems increased taxonomic and phylogenetic richness, diversity and heterogeneity of the soil microbiota when compared with conventional farming systems (Lupatini et al., 2017). In a simple definition, organic farming system consists of low-input agro-ecosystem farms in which plant productivity and ecosystem functionality are based on the natural availability of plant nutrients (Lammerts van Bueren et al., 2002). A study aimed at comparing the soil microbiome in conventional and organic farming systems in central Europe revealed no major differences among the main phyla of bacteria between the two farming styles (Armalytë et al., 2019), whereas another study that investigated the effects of 12 years of organic farming on soil microbiomes in northern China reported shifting of the community composition of dominant phyla and significant alterations of functional groups associated with ammonia oxidation, denitrification and phosphorus recycling when compared to conventional farming systems (Ding et al., 2019).
In addition to tillage, crop rotation also plays a pivotal role in increasing belowground microbial diversity compared to intensive mono-cropping practices. Although the United States Department of Agriculture has advocated [via the Conservation Reserve Program (CRP)] crop rotation to improve eroded land as early as 1985 (Allen and Vandever, 2005), its benefit on soil health has only been recognized recently. Several studies reported increases in such soil quality parameters as organic matter content, microbial biomass and respiration under crop rotation management when compared with a mono-cropping system (Campbell et al., 1991; Luce et al., 2013). A meta-analysis of 122 studies that examined crop rotation revealed similar findings, namely that adding one or more crops in rotation to a monoculture substantially increased the soil microbial biomass along with increases in total soil C and N, respectively (McDaniel et al., 2014). In another study, soil microbial communities of corn and switchgrass in mono-cropping systems when compared with mixed prairie grasses demonstrated that bacterial and fungal biomass, especially arbuscular mycorrhizal fungi, were higher in plots with mixed prairie grasses (Jesus et al., 2016). A 16S amplicon-based metagenomic analysis of an almost 20-year-old field trial in Bernburg, Germany revealed a significant effect of tillage practice and the preceding crop on prokaryotic community structures (Babin et al., 2019)
Cover crops are typically unharvested crops planted between cash crops that augment C provisioning to the soil system not only via unharvested residues, but also as root exudates that can support many rhizosphere microbes during the active growing season of the cover crop. Other benefits attributed to cover cropping include improved N fertility by incorporating legumes as a cover crop, reduced soil compaction via deep-rooted plants, and reduced erosion by keeping a plant and its root system in the field year round (Fernandez et al., 2016). Of various crop rotation management practices, those that include cover crops sustain soil quality and productivity by enhancing soil C, N and microbial biomass (Kim et al., 2020), making them a cornerstone for sustainable agroecosystems. Nonetheless, very few studies have assessed the relationship between cover crop stands and their associated belowground microbial communities. Early research in unfertilized grasslands demonstrated that fungal communities respond positively to plant-derived C inputs, suggesting that inclusion of cover crops in a rotation may promote fungal community development (Denef et al., 2009). More recently, a field study tested this hypothesis by specifically examining the impact on soil microbial communities of eight fall-sown cover crop species grown singly and in multispecies mixtures following a spring oats (Avena sativa L.) cropping season and found that certain cover crops selectively favored particular microbial functional groups. Arbuscular mycorrhizal fungi were more abundant beneath oat and cereal rye (Secale cereale L.) cover crops, while non-AM fungi were positively associated with hairy vetch (Vicia villosa L.) (Finney et al., 2017). Beyond positively affecting soil C and increasing the diversity of such beneficial fungi as arbuscular mycorrhiza, clover as a cover crop is often reported to suppress the relative abundance of pathogenic fungi (Benitez et al., 2016). Contrarily, in a 2-year field study, cover crops reportedly increased overall phylogenetic diversity of fungi but did not change the relative abundance of saprophytes, symbionts or pathogens, implying that cover cropping does not always appear to contribute to functional changes in the fungal community (Schmidt et al., 2019).
Reassessment of Plant Responsiveness to Symbiosis
It is now increasingly evident that plants employ fine-tuned mechanisms to shape the structure and function of their microbiome, with different genotypes of the same plant species growing in the same soil yet associating with distinct microbial communities (Berendsen et al., 2012). This is demonstrated in the findings of Bazghaleh et al. (2015), who clearly demonstrated the importance of intraspecific host variation in the association of chickpea cultivars with AM and non-AM fungi. Therefore, specific traits of a plant that modulate its microbiome should be considered as a trait for plant breeding (Wallenstein, 2017).
Despite the obvious importance of beneficial microorganisms for plant growth and fitness, and the impact of plant genotype on shaping their microbiome composition, plant germplasm is typically screened in the absence of microbes, and the selection of best breeding lines made solely based on the interaction between plant genotype and performance under various abiotic factors. We propose that an a priori examination of the interaction between a plant genotype(s) and the symbiotic microbes upon which it likely depends is an important factor in the selection of plant breeding lines. It seems very likely that a subset of rejected germplasm could outperform others, but only when coupled with a beneficial microbe or microbiome (Figure 2). Arguably, current breeding and selection efforts most likely result in decoupling of the soil microbiome from plant fitness. As a result, modern varieties may have lost their ability to support diverse microbiomes and thus, fail to gain the most from these interactions (Wallenstein, 2017).
It is now acknowledged that transitioning from a highly intensive mono-cropping system to a more diversified cropping system consisting of multiple host genotypes leads to increased bacterial and fungal diversity (Calderon et al., 2016). Hence, future plant breeding efforts should incorporate plant characteristics that are related to microbiome diversity. For example, efforts focusing on manipulating plant root exudates likely play a critical role in selective recruitment of the rhizosphere microbiome (Bakker et al., 2012). In support of this notion, it has been shown that plants can select which microbial populations receive the lion’s share of root exudates, demonstrating a capacity by the host to refine its microbial composition. Hence, an unbiased screening of plant genotypes for responsiveness in the presence of a beneficial microbe or microbiome can set forth a new and potentially transformative paradigm in selecting microbes for plant growth promotion (Figure 2).
Significance of Mycorrhizas: A Critical Component of Healthy Soil Rhizospheres
Mycorrhizae are mutualistic associations between soil fungi and plant roots that gradually evolved to be reciprocally beneficial to both partners (Brundrett, 2002). The benefits are generally assumed to involve an exchange of photosynthetically derived carbon from the host plant in exchange for soil nutrients provided by the foraging mycorrhiza. While likely true of arum-type arbuscular types of mycorrhizae, there are other types that can derive carbon from organic matter in the soil, or even “steal” it from one host plant to supply to another (Allen and Allen, 1991). A recent study has reported that in contrast to Arum maculatum, in which carbon is entirely derived from photo-assimilation, the green leaves of Paris quadrifolia contain a striking 50% carbon of fungal origin. Such partial mycoheterotrophy could thus potentially be widespread among the roughly 100,000 plant species that are known to develop a Paris-type AM, with far-reaching implications for our understanding of C trading in plant-microbe communities (Giesemann et al., 2019). Exactly what the mycorrhiza gains from this interaction is still under debate, but benefits may involve a safe haven from the open, more competitive soil space and a second, more reliable carbon source (Sapp, 2004).
Mycorrhizae not only shape plant communities, but they also affect the functional diversity of their cohabitants in the rhizospheric microbiome. The mycelium of mycorrhizal fungi transports plant-derived carbon into the soil in the form of sugars, amino acids and polyols to help sustain the microbiome (Tarkka et al., 2018). More recent studies focusing on soil microbial ecology revealed that mycorrhizal fungi mediate many diverse interactions within the soil “mycorrhizosphere,” including pathogens and mutualists that fix atmospheric nitrogen, take up phosphorus, produce vitamins, and/or protect against antagonists (Buée et al., 2009; Tedersoo et al., 2020). The “ectomycorrhizosphere,” which forms a very specific interface between soil and many trees, hosts a large and diverse community of microorganisms that likely play roles in mineral weathering and solubilization processes (Uroz et al., 2007). This carbon-rich mycorrhizosphere also supports large communities of root-associated microorganisms that further accelerate weathering of minerals by excreting organic acids, phenolic compounds, protons, and siderophores (Drever and Vance, 1994; Illmer et al., 1995).
Similarly, the extraradical hyphae of arbuscular mycorrhiza provide a direct pathway for the translocation of photosynthetically derived carbon to the soil, leading to the development of nutrient-rich niches for other soil microorganisms, particularly bacteria. A quantitative real-time PCR method detected significantly higher 16S rDNA abundance in both the bulk and the rhizosphere soils of zucchini (Cucurbita pepo L.) inoculated with Acaulospora laevis and Glomus mosseae (Qin et al., 2014). Additionally, arbuscular mycorrhizae have been reported to increase the relative abundance of Firmicutes, Streptomycetes, Comamonadaceae, and Oxalobacteraceae inhabiting the mycorrhizosphere (Offre et al., 2007; Nuccio et al., 2013). While there is clear evidence that microbial communities in the rhizosphere function cohesively with their mycorrhizal partner in nutrient mobilization from soil minerals, nitrogen cycling and protection of plants against root pathogens, such bidirectional synergy is not always universal. There are reports that indicate suppressive effects of bacterial communities on mycorrhizal functioning and vice versa. While one study reported (Svenningsen et al., 2018) that soil with a higher abundance of Acidobacteria suppresses the normal functioning of extra-radical mycelium in arbuscular mycorrhizae, another study found that Glomus intraradices and Glomus mosseae suppressed most of the associated soil microbial community (Welc et al., 2010).
A Novel Type of Endophytic Symbiont: The Serendipitaceae
A diverse group of fungi in the Basidiomycota, the Serendipitaceae (formerly Sebacinales Group B) (Oberwinkler et al., 2014) encompasses endophytes and lineages that repeatedly evolved ericoid, orchid and ectomycorrhizal types. Accordingly, in many natural ecosystems these fungi form mycorrhizal symbioses with an astounding variety of host plants – every mycorrhizal type, in fact, except for arbuscular. Previous research performed in our lab with a strain of this group, Serendipita vermifera, demonstrated plant growth-promoting properties in a variety of plants (Ghimire and Craven, 2011; Ray et al., 2015; Ray and Craven, 2016; Ray et al., 2020). Unfortunately, the agronomic utility of these fungi is hampered by the paucity of strains available, the large majority isolated from Australian orchids. We have begun to address this constraint by isolating the first North American strain of Serendipita, named Serendipita vermifera subsp. bescii NFPB0129, from the roots of a switchgrass plant in Ardmore, Oklahoma (Craven and Ray, 2017; Ray et al., 2018).
As mentioned above, soil organic matter has a tremendous influence on the biological, chemical, and physical properties of soils, making it a vital component of healthy agricultural systems. Whether a natural soil or an agricultural one, the release of the nutrients locked within SOM requires decomposers, primarily insects, fungi, and bacteria, to secrete organic acids and enzymes that can loosen and break down the cellulose and the recalcitrant lignin into nutritive forms that can be used by other microbes and plants. Unlike arbuscular mycorrhizae, which exchange inorganic, mineralized nutrients mined from the soil for carbon derived from host photosynthesis, members of the Serendipitaceae studied thus far have a complete arsenal of carbohydrate-active enzymes (CAZymes), representing approximately 4% of the entire gene set and rivaling the more well-studied saprophytic white and brown wood rotters, and much more than other symbiotic fungi. Additionally, genome analysis of S. bescii and S. vermifera suggests that Serendipitaceae fungi have the metabolic capacity to assimilate N from organic forms of N-containing compounds (Ray et al., 2019). We hypothesize that this carbohydrate-degrading enzyme complement endows these Serendipitaceae fungi with saprotrophic abilities (Craven and Ray, 2019). Unlike free-living decomposers that maintain a solitary lifestyle, seeking only dead or dying plant tissues as their source of subsistence, Serendipitaceae fungi seem to maintain a largely symbiotic lifestyle with the roots of living host plants. It currently is unclear whether there is expression of CAZymes while strains of Serendipita are in symbiosis with host plants, and if so, whether there is spatial or temporal separation from more mutualistic traits. Still, the capacity of some strains to form mycorrhizal relationships with orchids, where the seeds require carbon from the fungus for germination and often well into the plant’s lifespan, suggests that these Serendipitaceae symbionts may be less of a carbon cost to their host plant. Presumably, this saved carbon could potentially be used for other symbiotic relationships or developmental processes. In any case, these intriguing fungi and their seemingly unlimited host range provide a novel symbiosis that could be used in a broad variety of cropping systems.
Conclusion
Soil-dwelling microorganisms are critical components of soil health, itself a determinant of plant productivity and stress tolerance. Deploying microbes to improve agriculture productivity is an extremely attractive approach that is non-transgenic and can be viewed collectively as the extended plant genome. Because these same microbes can contribute to restoring soil health and productivity, they have a bright future in low-input, sustainable agriculture that extends beyond more classically defined plant-microbe symbioses.
Author Contributions
PR and KC conceived and planned the overall idea of the review manuscript. PR, VL, JL, and KC wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the Center for Bioenergy Innovation (CBI) project. The Center for Bioenergy Innovation (CBI) was a United States Department of Energy (DOE) Bioenergy Research Center supported by the Office of Biological and Environmental Research (OBER) in the DOE Office of Science.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors thank Josh Meo for graphic design.
References
Acosta-Martínez, V., Burow, G., Zobeck, T., and Allen, V. (2010). Soil microbial communities and function in alternative systems to continuous cotton. Soil Sci. Soc. Am. J. 74, 1181–1192. doi: 10.2136/sssaj2008.0065
Agler, M. T., Ruhe, J., Kroll, S., Morhenn, C., Kim, S.-T., Weigel, D., et al. (2016). Microbial hub taxa link host and abiotic factors to plant microbiome variation. PLoS Biol. 14:e1002352. doi: 10.1371/journal.pbio.1002352
Allen, A., and Vandever, M. (2005). The Conservation Reserve Program: planting for the future. US Geol. Surv. 2005:5145.
Allen, M. F., and Allen, M. F. (1991). The ecology of mycorrhizae. Cambridge: Cambridge University Press.
Armalytë, J., Skerniškytë, J., Bakienë, E., Krasauskas, R., Šiugždinienë, R., Kareivienë, V., et al. (2019). Microbial diversity and antimicrobial resistance profile in microbiota from soils of conventional and organic farming systems. Front. Microbiol. 10:892. doi: 10.3389/fmicb.2019.00892
Armanhi, J. S. L., de Souza, R. S. C., Damasceno, N. D. B., de Araújo, L. M., Imperial, J., and Arruda, P. (2018). A Community-Based Culture Collection for Targeting Novel Plant Growth-Promoting Bacteria from the Sugarcane Microbiome. Front. Plant Sci. 8:2191. doi: 10.3389/fpls.2017.02191
Arnold, A. E., Mejía, L. C., Kyllo, D., Rojas, E. I., Maynard, Z., Robbins, N., et al. (2003). Fungal endophytes limit pathogen damage in a tropical tree. Proc. Natl. Acad. Sci. 100, 15649–15654. doi: 10.1073/pnas.2533483100
Azad, K., and Kaminskyj, S. (2016). A fungal endophyte strategy for mitigating the effect of salt and drought stress on plant growth. Symbiosis 68, 73–78. doi: 10.1007/s13199-015-0370-y
Babin, D., Deubel, A., Jacquiod, S., Sørensen, S. J., Geistlinger, J., Grosch, R., et al. (2019). Impact of long-term agricultural management practices on soil prokaryotic communities. Soil Biol. Biochem. 129, 17–28. doi: 10.1016/j.soilbio.2018.11.002
Bais, H. P., Weir, T. L., Perry, L. G., Gilroy, S., and Vivanco, J. M. (2006). The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 57, 233–266. doi: 10.1146/annurev.arplant.57.032905.105159
Baker, K., and Cook, R. J. (1974). Biological control of plant pathogens. United states: WH Freeman and Company.
Bakker, M. G., Manter, D. K., Sheflin, A. M., Weir, T. L., and Vivanco, J. M. (2012). Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 360, 1–13. doi: 10.1007/s11104-012-1361-x
Banerjee, S., Schlaeppi, K., and van der Heijden, M. G. (2018). Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 16, 567–576. doi: 10.1038/s41579-018-0024-1
Banerjee, S., Walder, F., Büchi, L., Meyer, M., Held, A. Y., Gattinger, A., et al. (2019). Agricultural intensification reduces microbial network complexity and the abundance of keystone taxa in roots. ISME J. 13, 1722–1736. doi: 10.1038/s41396-019-0383-2
Baraza, E., Tauler, M., Romero-Munar, A., Cifre, J., and Gulias, J. (2016). “Mycorrhiza-based biofertilizer application to improve the quality of Arundo donax L., plantlets,” in Perennial Biomass Crops for a Resource-Constrained World, eds S. Barth, D. Murphy-Bokern, O. Kalinina, G. Taylor, and M. Jones (Cham: Springer), 225–232. doi: 10.1007/978-3-319-44530-4_19
Barelli, L., Waller, A. S., Behie, S. W., and Bidochka, M. J. (2020). Plant microbiome analysis after Metarhizium amendment reveals increases in abundance of plant growth-promoting organisms and maintenance of disease-suppressive soil. PLoS One 15:e0231150. doi: 10.1371/journal.pone.0231150
Bazghaleh, N., Hamel, C., Gan, Y., Tar’an, B., and Knight, J. D. (2015). Genotype-specific variation in the structure of root fungal communities is related to chickpea plant productivity. Appl. Environ. Microbiol. 81, 2368–2377. doi: 10.1128/aem.03692-14
Beirinckx, S., Viaene, T., Haegeman, A., Debode, J., Amery, F., Vandenabeele, S., et al. (2020). Tapping into the maize root microbiome to identify bacteria that promote growth under chilling conditions. Microbiome 8:54. doi: 10.1186/s40168-020-00833-w
Benitez, M.-S., Taheri, W. I., and Lehman, R. M. (2016). Selection of fungi by candidate cover crops. Appl. Soil Ecol. 103, 72–82. doi: 10.1016/j.apsoil.2016.03.016
Berendsen, R. L., Pieterse, C. M., and Bakker, P. A. (2012). The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486. doi: 10.1016/j.tplants.2012.04.001
Bergmann, J., Verbruggen, E., Heinze, J., Xiang, D., Chen, B., Joshi, J., et al. (2016). The interplay between soil structure, roots, and microbiota as a determinant of plant-soil feedback. Ecol. Evol. 6, 7633–7644. doi: 10.1002/ece3.2456
Bhattacharyya, P. N., and Jha, D. K. (2012). Plant growth-promoting rhizobacteria (PGPR): emergence in agriculture. World J. Microbiol. Biotechnol. 28, 1327–1350. doi: 10.1007/s11274-011-0979-9
Bodenhausen, N., Bortfeld-Miller, M., Ackermann, M., and Vorholt, J. A. (2014). A synthetic community approach reveals plant genotypes affecting the phyllosphere microbiota. PLoS Genet 10:e1004283. doi: 10.1371/journal.pgen.1004283
Bonfante, P., and Desirò, A. (2017). Who lives in a fungus? The diversity, origins and functions of fungal endobacteria living in Mucoromycota. ISME J. 11, 1727–1735. doi: 10.1038/ismej.2017.21
Bonfante, P., Venice, F., and Lanfranco, L. (2019). The mycobiota: fungi take their place between plants and bacteria. Curr. Opin. Microbiol. 49, 18–25. doi: 10.1016/j.mib.2019.08.004
Brundrett, M. C. (2002). Coevolution of roots and mycorrhizas of land plants. N. Phytol. 154, 275–304. doi: 10.1046/j.1469-8137.2002.00397.x
Brundrett, M. C., and Tedersoo, L. (2018). Evolutionary history of mycorrhizal symbioses and global host plant diversity. N. Phytol. 220, 1108–1115. doi: 10.1111/nph.14976
Buée, M., De Boer, W., Martin, F., Van Overbeek, L., and Jurkevitch, E. (2009). The rhizosphere zoo: an overview of plant-associated communities of microorganisms, including phages, bacteria, archaea, and fungi, and of some of their structuring factors. Plant Soil 321, 189–212. doi: 10.1007/s11104-009-9991-3
Bulgarelli, D., Schlaeppi, K., Spaepen, S., Van Themaat, E. V. L., and Schulze-Lefert, P. (2013). Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 64, 807–838. doi: 10.1146/annurev-arplant-050312-120106
Calderon, F. J., Nielsen, D., Acosta-Martinez, V., Vigil, M. F., and Drew, L. (2016). Cover crop and irrigation effects on soil microbial communities and enzymes in semiarid agroecosystems of the Central Great Plains of North America. Pedosphere 26, 192–205. doi: 10.1016/s1002-0160(15)60034-0
Calvo, P., Nelson, L., and Kloepper, J. W. (2014). Agricultural uses of plant biostimulants. Plant Soil 383, 3–41. doi: 10.1007/s11104-014-2131-8
Campbell, C., Biederbeck, V., Zentner, R., and Lafond, G. (1991). Effect of crop rotations and cultural practices on soil organic matter, microbial biomass and respiration in a thin Black Chernozem. Can. J. Soil Sci. 71, 363–376. doi: 10.4141/cjss91-035
Carlström, C. I., Field, C. M., Bortfeld-Miller, M., Müller, B., Sunagawa, S., and Vorholt, J. A. (2019). Synthetic microbiota reveal priority effects and keystone strains in the Arabidopsis phyllosphere. Nat. Ecol. Evol. 3, 1445–1454. doi: 10.1038/s41559-019-0994-z
Carrión, V. J., Perez-Jaramillo, J., Cordovez, V., Tracanna, V., De Hollander, M., Ruiz-Buck, D., et al. (2019). Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366, 606–612. doi: 10.1126/science.aaw9285
Castrillo, G., Teixeira, P. J. P. L., Paredes, S. H., Law, T. F., de Lorenzo, L., Feltcher, M. E., et al. (2017). Root microbiota drive direct integration of phosphate stress and immunity. Nature 543, 513–518. doi: 10.1038/nature21417
Chen, R., Senbayram, M., Blagodatsky, S., Myachina, O., Dittert, K., Lin, X., et al. (2014). Soil C and N availability determine the priming effect: microbial N mining and stoichiometric decomposition theories. Global Change Biol. 20, 2356–2367. doi: 10.1111/gcb.12475
Chutia, M., Mahanta, J., Bhattacheryya, N., Bhuyan, M., Boruah, P., and Sarma, T. (2007). Microbial herbicides for weed management: prospects, progress and constraints. Plant Pathol. J. 6, 210–218. doi: 10.3923/ppj.2007.210.218
Cordero, O. X., and Polz, M. F. (2014). Explaining microbial genomic diversity in light of evolutionary ecology. Nat. Rev. Microbiol. 12:263. doi: 10.1038/nrmicro3218
Craven, K. D., and Ray, P. (2019). More than Serendipity: The Potential to Manage Soil Carbon and Emissions While Promoting Low-Input Agriculture with Serendipitoid Mycorrhizae. Phytobio. J. 3, 161–164. doi: 10.1094/pbiomes-12-18-0058-p
Craven, K., and Ray, P. (2017). Symbiont for enhancement of plant performance. US Patent App. 1:926.
Dabral, S., Saxena, S. C., Choudhary, D. K., Bandyopadhyay, P., Sahoo, R. K., Tuteja, N., et al. (2020). Synergistic inoculation of Azotobacter vinelandii and Serendipita indica augmented rice growth. Cham: Springer.
del Barrio-Duque, A., Ley, J., Samad, A., Antonielli, L., Sessitsch, A., and Compant, S. (2019). Beneficial Endophytic Bacteria-Serendipita indica Interaction for Crop Enhancement and Resistance to Phytopathogens. Front. Microbiol. 10:2888. doi: 10.3389/fmicb.2019.02888
Denef, K., Roobroeck, D., Wadu, M. C. M., Lootens, P., and Boeckx, P. (2009). Microbial community composition and rhizodeposit-carbon assimilation in differently managed temperate grassland soils. Soil Biol. Biochem. 41, 144–153. doi: 10.1016/j.soilbio.2008.10.008
Ding, G.-C., Bai, M., Han, H., Li, H., Ding, X., Yang, H., et al. (2019). Microbial taxonomic, nitrogen cycling and phosphorus recycling community composition during long-term organic greenhouse farming. FEMS Microbiol. Ecol. 95:fiz042.
Donley, N. (2019). The USA lags behind other agricultural nations in banning harmful pesticides. Environ. Health 18:44. doi: 10.1186/s12940-019-0488-0
Drever, J. I., and Vance, G. F. (1994). “Role of soil organic acids in mineral weathering processes,” in Organic acids in geological processes, eds E. D. Pittman and M. D. Lewan (Berlin: Springer), 138–161. doi: 10.1007/978-3-642-78356-2_6
Eisenhauer, N., Lanoue, A., Strecker, T., Scheu, S., Steinauer, K., Thakur, M. P., et al. (2017). Root biomass and exudates link plant diversity with soil bacterial and fungal biomass. Scien. Rep. 7, 1–8.
Erktan, A., Cécillon, L., Graf, F., Roumet, C., Legout, C., and Rey, F. (2016). Increase in soil aggregate stability along a Mediterranean successional gradient in severely eroded gully bed ecosystems: combined effects of soil, root traits and plant community characteristics. Plant Soil 398, 121–137. doi: 10.1007/s11104-015-2647-6
Fatnassi, I. C., Chiboub, M., Saadani, O., Jebara, M., and Jebara, S. H. (2015). Impact of dual inoculation with Rhizobium and PGPR on growth and antioxidant status of Vicia faba L. under copper stress. Comp. Rend. Biol. 338, 241–254. doi: 10.1016/j.crvi.2015.02.001
Fernandez, A. L., Sheaffer, C. C., Wyse, D. L., Staley, C., Gould, T. J., and Sadowsky, M. J. (2016). Structure of bacterial communities in soil following cover crop and organic fertilizer incorporation. Appl. Microbiol. Biotechnol. 100, 9331–9341. doi: 10.1007/s00253-016-7736-9
Finkel, O. M., Castrillo, G., Paredes, S. H., González, I. S., and Dangl, J. L. (2017). Understanding and exploiting plant beneficial microbes. Curr. Opin. Plant Biol. 38, 155–163. doi: 10.1016/j.pbi.2017.04.018
Finney, D., Buyer, J., and Kaye, J. (2017). Living cover crops have immediate impacts on soil microbial community structure and function. J. Soil Water Conserv. 72, 361–373. doi: 10.2489/jswc.72.4.361
Frey-Klett, P., Garbaye, J. A., and Tarkka, M. (2007). The mycorrhiza helper bacteria revisited. N. Phytol. 176, 22–36. doi: 10.1111/j.1469-8137.2007.02191.x
Gao, Y., Yang, Y., Ling, W., Kong, H., and Zhu, X. (2011). Gradient Distribution of Root Exudates and Polycyclic Aromatic Hydrocarbons in Rhizosphere Soil. Soil Sci. Soc. Am. J. 75, 1694–1703. doi: 10.2136/sssaj2010.0244
Gerlagh, M. (1968). Introduction of Ophiobolus graminis into new polders and its decline. Netherlands J. Plant Pathol. 74, 1–97. doi: 10.1007/bf02019999
Ghimire, S. R., and Craven, K. D. (2011). Enhancement of switchgrass (Panicum virgatum L.) biomass production under drought conditions by the ectomycorrhizal fungus Sebacina vermifera. Appl. Environ, Microbiol. 77, 7063–7067. doi: 10.1128/AEM.05225-11
Giesemann, P., Rasmussen, H. N., Liebel, H. T., and Gebauer, G. (2019). Discreet heterotrophs: green plants that receive fungal carbon through Paris−type arbuscular mycorrhiza. N. Phytol. 226, 960–966. doi: 10.1111/nph.16367
Glick, B. R. (2012). Plant growth-promoting bacteria: mechanisms and applications. Scientifica 2012:963401.
Gunina, A., and Kuzyakov, Y. (2015). Sugars in soil and sweets for microorganisms: Review of origin, content, composition and fate. Soil Biol. Biochem. 90, 87–100. doi: 10.1016/j.soilbio.2015.07.021
Guo, H., Glaeser, S. P., Alabid, I., Imani, J., Haghighi, H., Kämpfer, P., et al. (2017). The abundance of endofungal bacterium Rhizobium radiobacter (syn. Agrobacterium tumefaciens) increases in its fungal host Piriformospora indica during the tripartite sebacinalean symbiosis with higher plants. Front. Microbiol. 8:629. doi: 10.3389/fmicb.2017.00629
Hartman, K., van der Heijden, M. G., Wittwer, R. A., Banerjee, S., Walser, J.-C., and Schlaeppi, K. (2018). Cropping practices manipulate abundance patterns of root and soil microbiome members paving the way to smart farming. Microbiome 6, 1–14. doi: 10.5810/kentucky/9780813175843.003.0001
Hartmann, A., Rothballer, M., and Schmid, M. (2008). Lorenz Hiltner, a pioneer in rhizosphere microbial ecology and soil bacteriology research. Plant Soil 312, 7–14. doi: 10.1007/s11104-007-9514-z
Hartmann, M., Frey, B., Mayer, J., Mäder, P., and Widmer, F. (2015). Distinct soil microbial diversity under long-term organic and conventional farming. ISME J. 9, 1177–1194. doi: 10.1038/ismej.2014.210
Hayat, R., Ali, S., Amara, U., Khalid, R., and Ahmed, I. (2010). Soil beneficial bacteria and their role in plant growth promotion: a review. Anna. Microbiol. 60, 579–598. doi: 10.1007/s13213-010-0117-1
Hayden, H. L., Savin, K. W., Wadeson, J., Gupta, V. V., and Mele, P. M. (2018). Comparative metatranscriptomics of wheat rhizosphere microbiomes in disease suppressive and non-suppressive soils for Rhizoctonia solani AG8. Front. Microbiol. 9:859. doi: 10.3389/fmicb.2018.00859
HoÈgberg, P., Nordgren, A., Buchmann, N., Taylor, A. F., Ekblad, A., HoÈgberg, M. N., et al. (2001). Large-scale forest girdling shows that current photosynthesis drives soil respiration. Nature 411, 789–792. doi: 10.1038/35081058
Hu, J., Wei, Z., Friman, V.-P., Gu, S.-H., Wang, X.-F., Eisenhauer, N., et al. (2016). Probiotic diversity enhances rhizosphere microbiome function and plant disease suppression. MBio 7, e1790–e1716.
Igiehon, N. O., and Babalola, O. O. (2017). Biofertilizers and sustainable agriculture: exploring arbuscular mycorrhizal fungi. Appl. Microbiol. Biotechnol. 101, 4871–4881. doi: 10.1007/s00253-017-8344-z
Illmer, P., Barbato, A., and Schinner, F. (1995). Solubilization of hardly-soluble AlPO4 with P-solubilizing microorganisms. Soil Biol. Biochem. 27, 265–270. doi: 10.1016/0038-0717(94)00205-f
Jeffries, P., and Rhodes, L. H. (1987). Use of mycorrhizae in agriculture. Crit. Rev. Biotechnol. 5, 319–357. doi: 10.3109/07388558709079476
Jesus, E. D. C., Liang, C., Quensen, J. F., Susilawati, E., Jackson, R. D., Balser, T. C., et al. (2016). Influence of corn, switchgrass, and prairie cropping systems on soil microbial communities in the upper Midwest of the United States. GCB Bioener. 8, 481–494. doi: 10.1111/gcbb.12289
Kemen, E. (2014). Microbe-microbe interactions determine oomycete and fungal host colonization. Curr. Opin. Plant Biol. 20, 75–81. doi: 10.1016/j.pbi.2014.04.005
Khatoon, Z., Huang, S., Rafique, M., Fakhar, A., Kamran, M. A., and Santoyo, G. (2020). Unlocking the potential of plant growth-promoting rhizobacteria on soil health and the sustainability of agricultural systems. J. Environ. Manag. 273:111118. doi: 10.1016/j.jenvman.2020.111118
Kim, N., Zabaloy, M. C., Guan, K., and Villamil, M. B. (2020). Do cover crops benefit soil microbiome? A meta-analysis of current research. Soil Biol. Biochem. 2020:107701. doi: 10.1016/j.soilbio.2019.107701
Klironomos, J. N., McCune, J., Hart, M., and Neville, J. (2000). The influence of arbuscular mycorrhizae on the relationship between plant diversity and productivity. Ecol. Lett. 3, 137–141. doi: 10.1046/j.1461-0248.2000.00131.x
Kumar, A., and Dubey, A. (2020). Rhizosphere microbiome: Engineering bacterial competitiveness for enhancing crop production. J. Adv. Res. 24, 337–352. doi: 10.1016/j.jare.2020.04.014
Kumar, U., Panneerselvam, P., Govindasamy, V., Vithalkumar, L., Senthilkumar, M., Banik, A., et al. (2017). Long-term aromatic rice cultivation effect on frequency and diversity of diazotrophs in its rhizosphere. Ecol. Engin. 101, 227–236. doi: 10.1016/j.ecoleng.2017.02.010
Kwak, M.-J., Kong, H. G., Choi, K., Kwon, S.-K., Song, J. Y., Lee, J., et al. (2018). Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 36, 1100–1109. doi: 10.1038/nbt.4232
Lakshmanan, V., Ray, P., and Craven, K. D. (2017). Toward a Resilient, Functional Microbiome: Drought Tolerance-Alleviating Microbes for Sustainable Agriculture. Plant Stress Tolerance 163, 69–84. doi: 10.1007/978-1-4939-7136-7_4
Lakshmanan, V., Selvaraj, G., and Bais, H. P. (2014). Functional soil microbiome: belowground solutions to an aboveground problem. Plant Physiol. 166, 689–700. doi: 10.1104/pp.114.245811
Lammerts van Bueren, E., Struik, P., and Jacobsen, E. (2002). Ecological concepts in organic farming and their consequences for an organic crop ideotype. Netherlands J. Agricult. Sci. 50, 1–26. doi: 10.1016/s1573-5214(02)80001-x
Lekberg, Y., and Helgason, T. (2018). In situ mycorrhizal function-knowledge gaps and future directions. N. Phytol. 220, 957–962. doi: 10.1111/nph.15064
Li, Y., Song, D., Liang, S., Dang, P., Qin, X., Liao, Y., et al. (2020a). Effect of no-tillage on soil bacterial and fungal community diversity: A meta-analysis. Soil Tillage Res. 204:104721. doi: 10.1016/j.still.2020.104721
Li, Y., Tremblay, J., Bainard, L. D., Cade−Menun, B., and Hamel, C. (2020b). Long−term effects of nitrogen and phosphorus fertilization on soil microbial community structure and function under continuous wheat production. Environ. Microbiol. 22, 1066–1088. doi: 10.1111/1462-2920.14824
Liu, Y., Zhu, A., Tan, H., Cao, L., and Zhang, R. (2019). Engineering banana endosphere microbiome to improve Fusarium wilt resistance in banana. Microbiome 7:74. doi: 10.1186/s40168-019-0690-x
Luce, M. S., Ziadi, N., Zebarth, B., Whalen, J., Grant, C., Gregorich, E., et al. (2013). Particulate organic matter and soil mineral nitrogen concentrations are good predictors of the soil nitrogen supply to canola following legume and non-legume crops in western Canada. Can. J. Soil Sci. 93, 607–620. doi: 10.4141/cjss2013-005
Luo, Y., Wang, F., Zhou, M., and Sheng, H. M. (2019). Sphingomonas sp. Cra20 increases plant growth rate and alters rhizosphere microbial community structure of Arabidopsis thaliana under drought stress. Front. Microbiol. 10:1221. doi: 10.3389/fmicb.2019.01221
Lupatini, M., Korthals, G. W., de Hollander, M., Janssens, T. K., and Kuramae, E. E. (2017). Soil microbiome is more heterogeneous in organic than in conventional farming system. Front. Microbiol. 7:2064. doi: 10.3389/fmicb.2016.02064
Marasco, R., Rolli, E., Ettoumi, B., Vigani, G., Mapelli, F., Borin, S., et al. (2012). A drought resistance-promoting microbiome is selected by root system under desert farming. PLloS One 7:e48479. doi: 10.1371/journal.pone.0048479
Marschner, P. (2012). Rhizosphere biology,” in Marschner’s Mineral Nutrition of Higher Plants (Third Edition). Netherland: Elsevier, 369–388.
Mathew, R. P., Feng, Y., Githinji, L., Ankumah, R., and Balkcom, K. S. (2012). Impact of no-tillage and conventional tillage systems on soil microbial communities. Appl. Environ. Soil Sci. 2012:548620.
Matson, P. A., Parton, W. J., Power, A. G., and Swift, M. J. (1997). Agricultural intensification and ecosystem properties. Science 277, 504–509. doi: 10.1126/science.277.5325.504
Mazzola, M. (2007). Manipulation of rhizosphere bacterial communities to induce suppressive soils. J. Nematol. 39:213.
Mbuthia, L. W., Acosta-Martínez, V., DeBruyn, J., Schaeffer, S., Tyler, D., Odoi, E., et al. (2015). Long term tillage, cover crop, and fertilization effects on microbial community structure, activity: Implications for soil quality. Soil Biol. Biochem. 89, 24–34. doi: 10.1016/j.soilbio.2015.06.016
McDaniel, M., Tiemann, L., and Grandy, A. (2014). Does agricultural crop diversity enhance soil microbial biomass and organic matter dynamics? A meta−analysis. Ecol. Applicat. 24, 560–570. doi: 10.1890/13-0616.1
McNear, D. H. Jr. (2013). The rhizosphere-roots, soil and everything in between. Nat. Educat. Knowl. 4:1.
Morella, N. M., Weng, F. C.-H., Joubert, P. M., Metcalf, C. J. E., Lindow, S., and Koskella, B. (2020). Successive passaging of a plant-associated microbiome reveals robust habitat and host genotype-dependent selection. Proc. Natl. Acad. Sci. 117, 1148–1159. doi: 10.1073/pnas.1908600116
Mueller, U. G., and Sachs, J. L. (2015). Engineering microbiomes to improve plant and animal health. Trends Microbiol. 23, 606–617. doi: 10.1016/j.tim.2015.07.009
Nadeem, S. M., Naveed, M., Zahir, Z. A., and Asghar, H. N. (2013). Plant microbe symbiosis: fundamentals and advances. Cham: Springer, 51–103.
Naito, M., Desirò, A., González, J. B., Tao, G., Morton, J. B., Bonfante, P., et al. (2017). ‘Candidatus Moeniiplasma glomeromycotorum’, an endobacterium of arbuscular mycorrhizal fungi. Int. J. of Syst. Evolut. Microbiol. 67, 1177–1184. doi: 10.1099/ijsem.0.001785
Nemergut, D. R., Schmidt, S. K., Fukami, T., O’Neill, S. P., Bilinski, T. M., Stanish, L. F., et al. (2013). Patterns and processes of microbial community assembly. Microbiol. Mol. Biol. Rev. 77, 342–356.
Nicot, P., Blum, B., Köhl, J., and Ruocco, M. (2011). Perspectives for future research-and-development projects on biological control of plant pests and diseases. Class. Augment. Biol. contr. Against Dis. Pests 2011, 68–70.
Nie, Y., Wang, M., Zhang, W., Ni, Z., Hashidoko, Y., and Shen, W. (2018). Ammonium nitrogen content is a dominant predictor of bacterial community composition in an acidic forest soil with exogenous nitrogen enrichment. Sci. Tot. Environ. 624, 407–415. doi: 10.1016/j.scitotenv.2017.12.142
Niu, B., Paulson, J. N., Zheng, X., and Kolter, R. (2017). Simplified and representative bacterial community of maize roots. Proc. Natl. Acad. Sci. U.S.A. 114, E2450–E2459. doi: 10.1073/pnas.1616148114
Nuccio, E. E., Hodge, A., Pett−Ridge, J., Herman, D. J., Weber, P. K., and Firestone, M. K. (2013). An arbuscular mycorrhizal fungus significantly modifies the soil bacterial community and nitrogen cycling during litter decomposition. Environ. Microbiol. 15, 1870–1881. doi: 10.1111/1462-2920.12081
Oberwinkler, F., Riess, K., Bauer, R., and Garnica, S. (2014). Morphology and molecules: the Sebacinales, a case study. Mycol. Progr. 13, 445–470. doi: 10.1007/s11557-014-0983-1
Offre, P., Pivato, B., Siblot, S., Gamalero, E., Corberand, T., Lemanceau, P., et al. (2007). Identification of bacterial groups preferentially associated with mycorrhizal roots of Medicago truncatula. Appl. Environ. Microbiol. 73, 913–921. doi: 10.1128/aem.02042-06
Oleńska, E., Małek, W., Wójcik, M., Swiecicka, I., Thijs, S., and Vangronsveld, J. (2020). Beneficial features of plant growth-promoting rhizobacteria for improving plant growth and health in challenging conditions: A methodical review. Sci. Tot. Environ. 32, 429–448.
Orwin, K. H., Wardle, D. A., and Greenfield, L. G. (2006). Ecological consequences of carbon substrate identity and diversity in a laboratory study. Ecology 87, 580–593. doi: 10.1890/05-0383
Panke-Buisse, K., Poole, A. C., Goodrich, J. K., Ley, R. E., and Kao-Kniffin, J. (2014). Selection on soil microbiomes reveals reproducible impacts on plant function. ISME J. 9:980. doi: 10.1038/ismej.2014.196
Paredes, S. H., Gao, T., Law, T. F., Finkel, O. M., Mucyn, T., Teixeira, P. J. P. L., et al. (2018). Design of synthetic bacterial communities for predictable plant phenotypes. PLoS biology 16:e2003962. doi: 10.1371/journal.pbio.2003962
Parnell, J. J., Berka, R., Young, H. A., Sturino, J. M., Kang, Y., Barnhart, D., et al. (2016). From the lab to the farm: an industrial perspective of plant beneficial microorganisms. Front. Plant Sci. 7:1110. doi: 10.3389/fpls.2016.01110
Paul, K., Saha, C., Nag, M., Mandal, D., Naiya, H., Sen, D., et al. (2020). A tripartite interaction among the basidiomycete Rhodotorula mucilaginosa, N2-fixing endobacteria, and rice improves plant nitrogen nutrition. Plant Cell 32, 486–507. doi: 10.1105/tpc.19.00385
Priyadharsini, P., and Muthukumar, T. (2016). “Interactions between arbuscular mycorrhizal fungi and potassium-solubilizing microorganisms on agricultural productivity,” in Potassium solubilizing microorganisms for sustainable agriculture, eds V. S. Meena, B. R. Maurya, and J. P. Verma (Cham: Springer), 111–125. doi: 10.1007/978-81-322-2776-2_8
Qin, H., Brookes, P. C., Xu, J., and Feng, Y. (2014). Bacterial degradation of Aroclor 1242 in the mycorrhizosphere soils of zucchini (Cucurbita pepo L.) inoculated with arbuscular mycorrhizal fungi. Environ. Sci. Pollut. Res. 21, 12790–12799. doi: 10.1007/s11356-014-3231-y
Rai, P. K., Singh, M., Anand, K., Saurabh, S., Kaur, T., Kour, D., et al. (2020). New and Future Developments in Microbial Biotechnology and Bioengineering, Netherland: Elsevier, 49–60.
Raupp, J., Pekrun, C., Oltmanns, M., and Köpke, U. (2006). “The Rodale farming system trial 1981–2005: long term analysis of organic and conventional maize and soybean cropping systems,”in Long-term field experiments in organic farming, eds J. Raupp, C. Pekrun, M. Oltmanns, and U. Köpke (Germany: Institute for Biodynamic Research).
Ray, P., Abraham, P. E., Guo, Y., Giannone, R. J., Engle, N. L., Yang, Z. K., et al. (2019). Scavenging organic nitrogen and remodeling lipid metabolism are key survival strategies adopted by the endophytic fungi, Serendipita vermifera and Serendipita bescii to alleviate nitrogen and phosphorous starvation in vitro. Environ. Microbiol. Rep. 11, 548–557. doi: 10.1111/1758-2229.12757
Ray, P., and Craven, K. D. (2016). Sebacina vermifera: a unique root symbiont with vast agronomic potential. World J. Microbiol. Biotechnol. 32, 1–10.
Ray, P., Chi, M.-H., Guo, Y., Chen, C., Adam, C., Kuo, A., et al. (2018). Genome sequence of the plant growth-promoting fungus Serendipita vermifera subsp. bescii: The first native strain from North America. Phytobiomes 2, 2471–2906.
Ray, P., Guo, Y., Chi, M.-H., Krom, N., Saha, M. C., and Craven, K. D. (2020). Serendipita bescii promotes winter wheat growth and modulates the host root transcriptome under phosphorus and nitrogen starvation. Environ. Microbiol. 2020:15242. doi: 10.1111/1462-2920.15242
Ray, P., Ishiga, T., Decker, S. R., Turner, G. B., and Craven, K. D. (2015). A Novel Delivery System for the Root Symbiotic Fungus, Sebacina vermifera, and Consequent Biomass Enhancement of Low Lignin COMT Switchgrass Lines. BioEner. Res. 8, 922–933. doi: 10.1007/s12155-015-9636-8
Rogers, C., and Oldroyd, G. E. (2014). Synthetic biology approaches to engineering the nitrogen symbiosis in cereals. J. Exp. Bot. 65, 1939–1946. doi: 10.1093/jxb/eru098
Ryu, M.-H., Zhang, J., Toth, T., Khokhani, D., Geddes, B. A., Mus, F., et al. (2020). Control of nitrogen fixation in bacteria that associate with cereals. Nat. Microbiol. 5, 314–330. doi: 10.1038/s41564-019-0631-2
Saad, M. M., Eida, A. A., and Hirt, H. (2020). Tailoring plant-associated microbial inoculants in agriculture: a roadmap for successful application. J. Exp. Bot. 71, 3878–3901. doi: 10.1093/jxb/eraa111
Sapp, J. (2004). The dynamics of symbiosis: an historical overview. Can. J. Bot. 82, 1046–1056. doi: 10.1139/b04-055
Sarkar, A., Pramanik, K., Mitra, S., Soren, T., and Maiti, T. K. (2018). Enhancement of growth and salt tolerance of rice seedlings by ACC deaminase-producing Burkholderia sp. MTCC 12259. J. Plant Physiol. 231, 434–442. doi: 10.1016/j.jplph.2018.10.010
Schmid, C. A., Schröder, P., Armbruster, M., and Schloter, M. (2018). Organic amendments in a long-term field trial—consequences for the bulk soil bacterial community as revealed by network analysis. Microbial. Ecol. 76, 226–239. doi: 10.1007/s00248-017-1110-z
Schmidt, R., Mitchell, J., and Scow, K. (2019). Cover cropping and no-till increase diversity and symbiotroph: saprotroph ratios of soil fungal communities. Soil Biol. Biochem. 129, 99–109. doi: 10.1016/j.soilbio.2018.11.010
Schneider, A. (1892). Observations on Some American Rhizobia. Bull. Torrey Botan. Club 19, 203–218. doi: 10.2307/2475812
Schreiter, S., Sandmann, M., Smalla, K., and Grosch, R. (2014). Soil type dependent rhizosphere competence and biocontrol of two bacterial inoculant strains and their effects on the rhizosphere microbial community of field-grown lettuce. PLoS One 9:e103726. doi: 10.1371/journal.pone.0103726
Schweitzer, J. A., Bailey, J. K., Fischer, D. G., LeRoy, C. J., Lonsdorf, E. V., Whitham, T. G., et al. (2008). Plant-soil-microorganism interactions: Heritable relationship between plant genotype and associated soil microorganisms. Ecology 89, 773–781. doi: 10.1890/07-0337.1
Sergaki, C., Lagunas, B., Lidbury, I., Gifford, M. L., and Schäfer, P. (2018). Challenges and approaches in microbiome research: from fundamental to applied. Front. Plant Sci. 9:1205. doi: 10.3389/fpls.2018.01205
Shipton, P., Cook, R., and Sitton, J. (1973). Occurrence and Transfer of a Biological. Phytopathology 63, 511–517. doi: 10.1094/phyto-63-511
Singh, D., Ghosh, P., Kumar, J., and Kumar, A. (2019). Microbial Interventions in Agriculture and Environment. Cham: Springer, 205–227.
Singh, S. (2016). Microbial Inoculants in Sustainable Agricultural Productivity. Cham: Springer, 69–83.
Souza, R. D., Ambrosini, A., and Passaglia, L. M. (2015). Plant growth-promoting bacteria as inoculants in agricultural soils. Genet. Mole. Biol. 38, 401–419. doi: 10.1590/s1415-475738420150053
Sun, C., Johnson, J. M., Cai, D., Sherameti, I., Oelmüller, R., and Lou, B. (2010). Piriformospora indica confers drought tolerance in Chinese cabbage leaves by stimulating antioxidant enzymes, the expression of drought-related genes and the plastid-localized CAS protein. J. Plant Physiol. 167, 1009–1017. doi: 10.1016/j.jplph.2010.02.013
Sun, R., Zhang, X.-X., Guo, X., Wang, D., and Chu, H. (2015). Bacterial diversity in soils subjected to long-term chemical fertilization can be more stably maintained with the addition of livestock manure than wheat straw. Soil Biol. Biochem. 88, 9–18. doi: 10.1016/j.soilbio.2015.05.007
Svenningsen, N. B., Watts-Williams, S. J., Joner, E. J., Battini, F., Efthymiou, A., Cruz-Paredes, C., et al. (2018). Suppression of the activity of arbuscular mycorrhizal fungi by the soil microbiota. ISME J. 12:1296.
Tarkka, M. T., Drigo, B., and Deveau, A. (2018). Mycorrhizal microbiomes. Mycorrhiza 28, 403–409. doi: 10.1007/s00572-018-0865-5
Tedersoo, L., Bahram, M., and Zobel, M. (2020). How mycorrhizal associations drive plant population and community biology. Science 367: 6480.
Tian, J., He, N., Hale, L., Niu, S., Yu, G., Liu, Y., et al. (2018). Soil organic matter availability and climate drive latitudinal patterns in bacterial diversity from tropical to cold temperate forests. Funct. Ecol. 32, 61–70. doi: 10.1111/1365-2435.12952
Timmusk, S., Behers, L., Muthoni, J., Muraya, A., and Aronsson, A.-C. (2017). Perspectives and challenges of microbial application for crop improvement. Front. Plant Sci. 8:49. doi: 10.3389/fpls.2017.00049
Toyota, K., and Shirai, S. (2018). Growing interest in microbiome research unraveling disease suppressive soils against plant pathogens. Microbes Environ. 33, 345–347. doi: 10.1264/jsme2.me3304rh
Tsolakidou, M.-D., Stringlis, I. A., Fanega-Sleziak, N., Papageorgiou, S., Tsalakou, A., Pantelides, I. S., et al. (2019). Rhizosphere-enriched microbes as a pool to design synthetic communities for reproducible beneficial outputs. FEMS Microbiol. Ecol. 95:fiz138. doi: 10.1093/femsec/fiz138
Turrini, A., Avio, L., Giovannetti, M., and Agnolucci, M. (2018). Functional complementarity of arbuscular mycorrhizal fungi and associated microbiota: the challenge of translational research. Front. Plant Sci. 9:1407. doi: 10.3389/fpls.2018.01407
Uroz, S., Calvaruso, C., Turpault, M.-P., Pierrat, J.-C., Mustin, C., and Frey-Klett, P. (2007). Effect of the mycorrhizosphere on the genotypic and metabolic diversity of the bacterial communities involved in mineral weathering in a forest soil. Appl. Environ. Microbiol. 73, 3019–3027. doi: 10.1128/aem.00121-07
Vandenkoornhuyse, P., Quaiser, A., Duhamel, M., Le Van, A., and Dufresne, A. (2015). The importance of the microbiome of the plant holobiont. N. Phytol. 206, 1196–1206. doi: 10.1111/nph.13312
Verbruggen, E., Van Der Heijden, M. G., Weedon, J. T., Kowalchuk, G. A., and Röling, W. F. (2012). Community assembly, species richness and nestedness of arbuscular mycorrhizal fungi in agricultural soils. Mole. Ecol. 21, 2341–2353. doi: 10.1111/j.1365-294x.2012.05534.x
Vorholt, J. A., Vogel, C., Carlström, C. I., and Müller, D. B. (2017). Establishing causality: opportunities of synthetic communities for plant microbiome research. Cell Host Microbe 22, 142–155. doi: 10.1016/j.chom.2017.07.004
Wallenstein, M. D. (2017). Managing and manipulating the rhizosphere microbiome for plant health: a systems approach. Rhizosphere 3, 230–232. doi: 10.1016/j.rhisph.2017.04.004
Wang, F., Chen, S., Wang, Y., Zhang, Y., Hu, C., and Liu, B. (2018). Long-term nitrogen fertilization elevates the activity and abundance of nitrifying and denitrifying microbial communities in an upland soil: implications for nitrogen loss from intensive agricultural systems. Front. Microbiol. 9:2424. doi: 10.3389/fmicb.2018.02424
Welc, M., Ravnskov, S., Kieliszewska-Rokicka, B., and Larsen, J. (2010). Suppression of other soil microorganisms by mycelium of arbuscular mycorrhizal fungi in root-free soil. Soil Biol. Biochem. 42, 1534–1540. doi: 10.1016/j.soilbio.2010.05.024
Weller, D. M., Raaijmakers, J. M., Gardener, B. B. M., and Thomashow, L. S. (2002). Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu. Rev. Phytopathol. 40, 309–348.
Weyens, N., van der Lelie, D., Taghavi, S., Newman, L., and Vangronsveld, J. (2009). Exploiting plant-microbe partnerships to improve biomass production and remediation. Trends Biotechnol. 27, 591–598. doi: 10.1016/j.tibtech.2009.07.006
Wild, B., Schnecker, J., Alves, R. J. E., Barsukov, P., Bárta, J., Èapek, P., et al. (2014). Input of easily available organic C and N stimulates microbial decomposition of soil organic matter in arctic permafrost soil. Soil Biol. Biochem. 75, 143–151. doi: 10.1016/j.soilbio.2014.04.014
Yadav, R., Ror, P., Rathore, P., and Ramakrishna, W. (2020). Bacteria from native soil in combination with arbuscular mycorrhizal fungi augment wheat yield and biofortification. Plant Physiol. Biochem. 150, 222–233. doi: 10.1016/j.plaphy.2020.02.039
Yergeau, E., Bell, T. H., Champagne, J., Maynard, C., Tardif, S., Tremblay, J., et al. (2015). Transplanting soil microbiomes leads to lasting effects on willow growth, but not on the rhizosphere microbiome. Front. Microbiol. 6:1436. doi: 10.3389/fmicb.2015.01436
Yin, C., Mueth, N., Hulbert, S., Schlatter, D., Paulitz, T. C., Schroeder, K., et al. (2017). Bacterial communities on wheat grown under long-term conventional tillage and no-till in the Pacific Northwest of the United States. Phytobiomes 1, 83–90. doi: 10.1094/pbiomes-09-16-0008-r
Yuan, Z. L., Druzhinina, I. S., Labbé, J., Redman, R. S., Qin, Y., Rodriguez, R., et al. (2016). Specialized microbiome of a halophyte and its role in helping non-host plants to withstand salinity. Sci. Rep. 6:32467. doi: 10.1038/srep32467
Hotspot Quiz
Check your Understanding
- What roles does the soil microbiome have in the movement towards sustainable agriculture?
- What is a rhizosphere?
- What type of soil fungi are associated with promoting plant health as well as the soil microbiome? How do they do so?
- What are the challenges with studying plant growth promoting microorganisms?
- Explain the difference between a reductionist vs. holistic approach for improving plant growth with microorganisms.
- What types of agricultural practices can alter the soil microbiome?
- How could the use of ‘probiotics’ suppress disease-causing soil microorganisms?
Media Attributions
- Video 1 – Understanding Our Soil: The Nitrogen Cycle, Fixers, and Fertilizer by Jimi Sol. Licensed under Creative Commons: By Attribution 3.0 License https://creativecommons.org/licenses/by/3.0/
- Hot Spot Quiz Figure – Plant and Soil Microbiomes by Gopal & Gupta, 2016 licensed under Creative Commons Attribution-Share Alike 4.0
References
- Gopal, M., & Gupta, A. (2016). Microbiome Selection Could Spur Next-Generation Plant Breeding Strategies. Frontiers in microbiology, 7, 1971. https://doi.org/10.3389/fmicb.2016.01971
-
Omotayo, O. P., & Babalola, O. O. (2021). Resident rhizosphere microbiome’s ecological dynamics and conservation: Towards achieving the envisioned Sustainable Development Goals, a review. International Soil and Water Conservation Research, 9(1), 127–142. https://doi.org/10.1016/j.iswcr.2020.08.002
-
Ray, P., Lakshmanan, V., Labbé, J. L., & Craven, K. D. (2020). Microbe to Microbiome: A Paradigm Shift in the Application of Microorganisms for Sustainable Agriculture. Frontiers in Microbiology, 11. https://www.frontiersin.org/article/10.3389/fmicb.2020.622926
- Tosi, M., Mitter, E. K., Gaiero, J., & Dunfield, K. (2020). It takes three to tango: the importance of microbes, host plant, and soil management to elucidate manipulation strategies for the plant microbiome. Canadian Journal of Microbiology, 66(7), 413–433. https://doi.org/10.1139/cjm-2020-0085

